
Practice Perfect 793
Diagnosing MRSA From a Sensitivity Report...
Could You Answer This Residency Interview Question?
Diagnosing MRSA From a Sensitivity Report...
Could You Answer This Residency Interview Question?
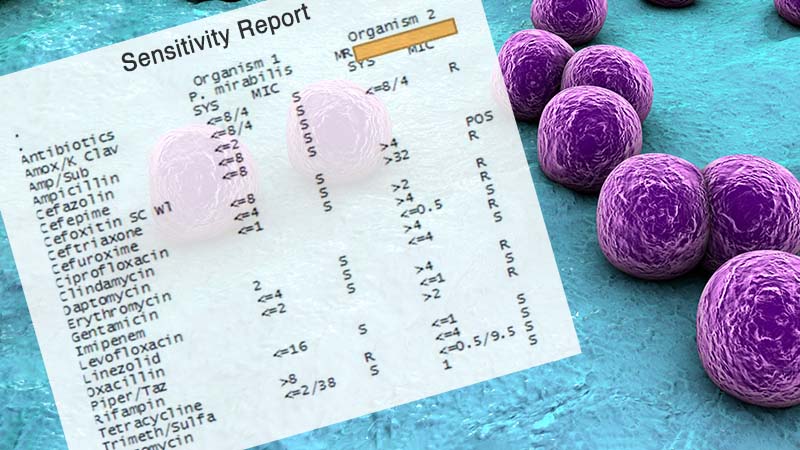
Every year, during the Chino Valley Medical Center residency interviews, there appears to be one common topic that stumps all candidates, regardless from what school they come. Last year, it was fundamental biomechanics (I asked the simple question, “Name two foot orthotic modifications for a patient with heel pain and post-static dyskinesia”). Yes, last year, students had a hard time answering that question. Painful, I know. I’m happy to report this year was much better from a biomechanical standpoint.
The painfully, unknown topic this year? Infectious disease, specifically how to diagnose methicillin resistant Staphylococcus aureus (MRSA) from a sensitivity report. Since I won’t use this question again during future interviews, let’s go through the question and discuss the answer.
This was the slide we presented. It was part of a larger case work-up of a diabetic foot infection:
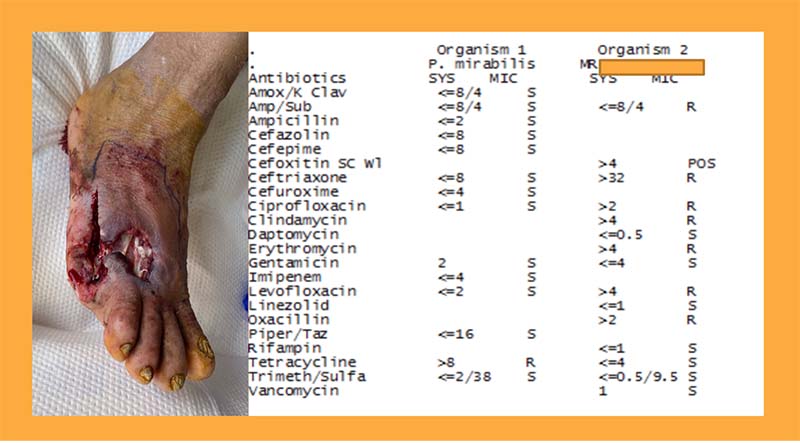
The question we asked was: “This is the appearance on postop day 3 and the culture and sensitivity. One of the organisms was Proteus mirabilis. Can you tell us what the other organism was? What antibiotics would you choose to treat this?”
First of all, the image of the foot is a gruesome distractor that makes absolutely no difference for the case. It’s just there for continuity with the rest of the case (and makes the interview harder). The answers, though, were somewhat incongruent. I received answers including vancomycin-resistant enterococcus (not possible since the organism is vancomycin sensitive) and methicillin-sensitive Staphylococcus aureus (again, not possible which I’ll explain in a second), among others. Students explained their answers by stating drugs to which the organism was sensitive. Good try, but wrong.
Here’s your answer: Organism 2 is resistant to…Oxacillin so the correct answer is MRSA. That’s it. That’s the answer. Methicillin resistance is tested and reported using oxacillin. I know – it’s kind of anticlimactic and possibly confusing at the same time. Let’s review how this is possible, including some of the fundamental concepts our students should have known (and some other fun facts!).
Here’s the answer: Organism 2 is resistant to oxacillin, so the correct answer is…MRSA. Methicillin resistance is tested and reported using oxacillin.
MRSA infection is defined by the United States Clinical Laboratory Standards Institute (CLSI) as having an oxacillin minimum inhibitory concentration (MIC) of > ≥4 mcg/mL.1 The European version of this organization defines it as >2 mg/L,2 which is what my local hospital apparently uses as its cutoff. It is also possible (and may even be better) to use cefoxitin to determine MRSA status, and as you can see in the case, the MIC for cefoxitin was > 4 and states “POS”. Hint hint, students!
The resistance for MRSA we are concerned about is that to beta-lactam antibiotics, including penicillins such as oxacillin and methicillin, and cephalosporins. This resistance comes from the acquisition of the mecA gene by a Staphylococcus aureus bacterium. This gene encodes for the penicillin binding protein 2a (PBP 2a) which prevents penicillins and cephalosporins from binding to the peptidoglycan layer of the bacterial cell wall, allowing it to continue to live.
OMG!!! Flashbacks to medical school. Stop, Shapiro, please!!!!!
Ok, Ok. Take a deep breath and relax. Moving on. Sort of…
I can’t help but report the results of an interesting recent paper in Nature by Larsen, et al that found strains of MRSA actually predate the antibiotic era.3 It wasn’t just our overuse of antibiotics that lead to MRSA. It has actually been found in European hedgehogs, entering the human population as a result of our common interaction with these animals (in addition to our overuse of antibiotics). These researchers also found that Trichophyton erinaceid, a dermatophyte fungus living on the skin of these animals, produces two beta-lactam antibiotics, creating an evolutionary pressure that favors those staph bacteria that produce resistance genes and, hence, MRSA. That brings a new meaning to the “Sonic Hedgehog” game.
But wait, there’s more. Hang on, friends, we’re going down the rabbit hole!!!
Now, I’ll bet you may not have heard of Trichophyton erinaceid, but “Trichophyton” sure sounds familiar, right? As in Trichophyton rubrum and Trichophyton mentagrophytes, two common causes of tinea infections, including onychomycosis and tinea pedis. If human Trichophyton species produced antibiotics, then onychomycosis and tinea pedis could conceivably be breeding grounds for MRSA bacteria.
And, in fact, Hammadi and colleagues, found in 2007 that some species of Trichophyton, including T. mentagrophytes, produce antibiotic-like substances!4 Wow, could there be a link between onychomycosis and MRSA infections? Could we be creating our own evolutionary pressures leading to antibacterial resistant organisms by undertreating dermatophytosis and onychomycosis? Is it also possible that onychomycosis organisms could harbor the next great antimicrobial medication? I’ll leave that up to better minds than my own and return to MRSA.
Wow, could there be a link between onychomycosis and MRSA infections?
Climbing back out of that rabbit hole (phew, we made it!), let’s answer that final question about why oxacillin is used to determine MRSA status rather than methicillin. In 1959 methicillin was introduced as the first semisynthetic penicillinase-resistant penicillin, and in 1961, when MRSA was first found, it would have made sense to use methicillin. However, methicillin was pulled from the market due to cases of interstitial nephritis. It was found that oxacillin, a derivative of methicillin, approved for medical use in 1962, did not cause the same problems as methicillin. It was also more shelf stable and more likely to detect resistant strains. Staph aureus strains are, then, resistant to both methicillin and oxacillin, so MRSA is determined using the better oxacillin. Why don’t we use ORSA instead of MRSA? The name just stuck. Besides, why should we make it so easy for our podiatry residency applicants?
Best wishes.

Jarrod Shapiro, DPM
PRESENT Practice Perfect Editor
[email protected]

-
National Committee for Clinical Laboratory Standards. Performance standards for antimicrobial susceptibility testing. CLSI approved standard M100-S23. Clinical and Laboratory Standards Institute, Wayne, PA 2013
-
European Committee on Antimicrobial Susceptibility Testing. Breakpoint tables for interpretation of MICs and zone diameters. Version 8.1, 2018. www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/Breakpoint_tables/v_8.1_Breakpoint_Tables.pdf (Accessed on August 09, 2018).
Follow this link -
Larsen J, Raisen CL, Ba X, Sadgrove NJ, Padilla-González GF, Simmonds MS, Loncaric I, Kerschner H, Apfalter P, Hartl R, Deplano A. Emergence of methicillin resistance predates the clinical use of antibiotics. Nature. 2022 Jan 5:1-7.
Follow this link -
Hammadi DK, Selselet GA, Bensoultane SA. Antibiotic like-substances produced by some trichophytic dermatophytes. Afr J Biotechnol. 2007;6(15):1788-1790.
Follow this link
































Comments
There are 0 comments for this article